This is the most shocking thing I have seen yet regarding the COVID vaccines. The SV40 promoter is a means to move plasmid DNA into cell nuclei.
Has the human race just lived through a deliberate experiment to disrupt and change its genome? h/t Sasha Latypova (again)
Here is Sasha’s more cmprehensive article of Sept. 1:
What if that huge amount of E.coli-made double-stranded DNA encased in plasmids was in the mRNA COVID vaccines for a reason, and was not the result of sloppy manufacturing?
What if it was there to make it interact with our own genes, with the intention to cause mutations in our chromosomes?
And what if the SV40 promoter recently found in the vaccine was there deliberately, to enhance this process, since it has a unique function of allowing the plasmid DNA to get through the nuclear membrane, to enter the nucleus? There are a lot of “What ifs” here.
My laboratory studies the mechanisms and applications of plasmid and DNA-binding protein nuclear localization. Our long term goals are to develop gene therapy approaches to the treatment of a variety of human diseases by focusing on the development of novel non-viral intra- and extracellular delivery methods. Our main emphasis is in the area of pulmonary gene delivery and function. Perhaps the major problem hindering gene therapy is the inefficiency of gene transfer to slowly and non-dividing cells. While many aspects of non-viral vector design are being addressed, one critical area that has not received adequate attention is the nuclear import of vector DNA. Clearly, without the translocation of plasmid DNA into the nucleus, no gene expression, or "gene therapy" can take place. My laboratory continues to identify and characterize novel DNA sequences to promote nuclear import of non-viral vectors, both in cultured cells and in vivo, as well as sequences that promote cytoplasmic and intranuclear trafficking.
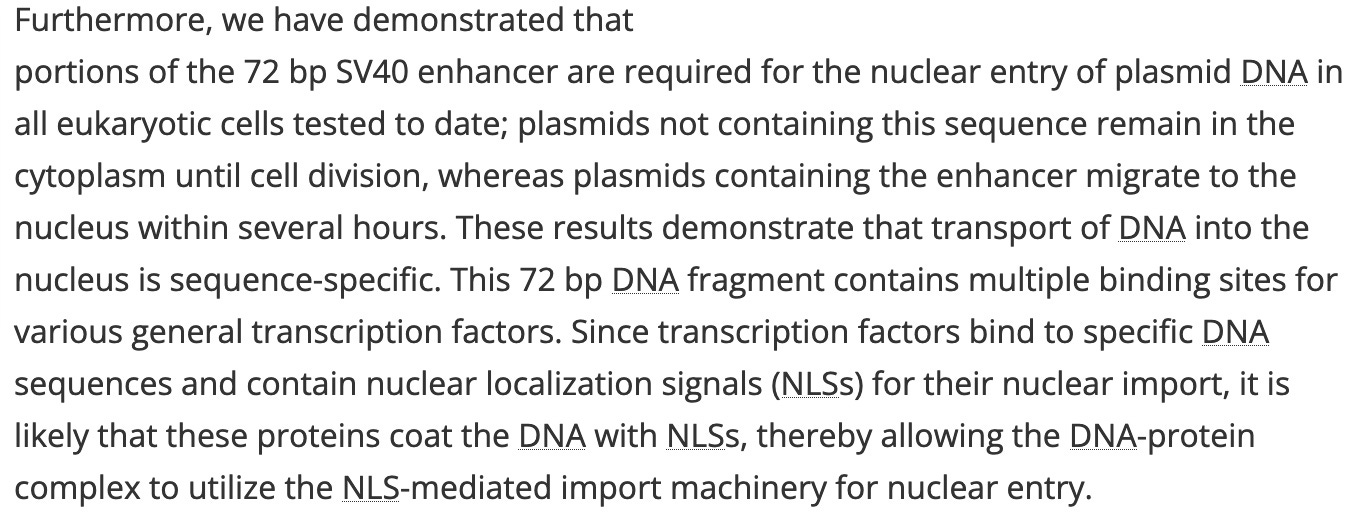
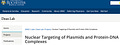
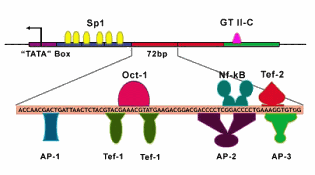
Organization of transcription factor binding sites on the SV40 enhancer.
Over the past 15 years, work from our laboratory has addressed the nuclear targeting and entry of plasmid DNA. Using cultured cells, we have shown that plasmids are able to enter the nuclei of cells in the absence of cell division and its accompanying nuclear envelope breakdown. Assays used to follow the movement of DNA include in situ hybridization, reporter gene expression, GFP-, YFP-, and RFP- tagged proteins, and live cell imaging of fluorescently-labeled plasmids and RNAs. As for all other macromolecular exchange between the cytoplasm and nucleus, DNA nuclear entry is mediated by the nuclear pore complex.
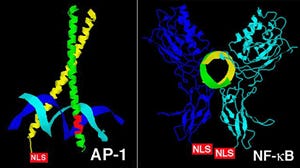
Crystal structures of AP1 (left) and NF-kB (right) transcription factors. Note that when bound to DNA, the NLS's of both proteins are likely accessible to import proteins.
Furthermore, we have demonstrated that portions of the 72 bp SV40 enhancer are required for the nuclear entry of plasmid DNA in all eukaryotic cells tested to date; plasmids not containing this sequence remain in the cytoplasm until cell division, whereas plasmids containing the enhancer migrate to the nucleus within several hours. These results demonstrate that transport of DNA into the nucleus is sequence-specific. This 72 bp DNA fragment contains multiple binding sites for various general transcription factors. Since transcription factors bind to specific DNA sequences and contain nuclear localization signals (NLSs) for their nuclear import, it is likely that these proteins coat the DNA with NLSs, thereby allowing the DNA-protein complex to utilize the NLS-mediated import machinery for nuclear entry.
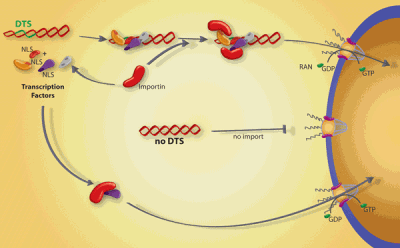
Model for sequence-specific plasmid nuclear import.
Interestingly, the SV40 sequence appears to be somewhat unique among viral enhancers: neither the strong promoter/enhancers of CMV or RSV have similar DNA nuclear import activity. More recent work from our laboratory has taken a proteomics approach to identify proteins and protein complexes needed for this nuclear targeting and import. We have shown that not only are key transcription factors involved in the nuclear import of plasmids, but that NLS-receptor proteins, chaperones, and a host of other proteins also play a vitally important role in nuclear localization.
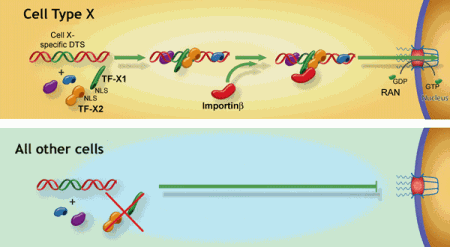
Model for cell-specific plasmid nuclear import.
Our model predicts that by using DNA elements containing binding sites for transcription factors expressed in unique cell types, we should be able to create plasmids that target to the nucleus in a cell-specific manner. Using the promoter from the smooth muscle gamma actin (SMGA) gene whose expression is limited to smooth muscle cells, we have created a series of reporter plasmids that are expressed selectively in smooth muscle cells. Moreover, when injected into the cytoplasm or delivered to cells by liposome-, polyplex-, or electroporation-mediated transfection, plasmids containing portions of the SMGA promoter localize to the nucleus of smooth muscle cells, but remain cytoplasmic in fibroblasts, epithelial cells, and endothelial cells. In contrast, a similar plasmid carrying the SV40 enhancer is transported into the nuclei of all cell types tested. Using a mutagenesis approach, we found that the presence of binding sites for two transcription factors, SRF and Nkx3, is needed for this cell-specific nuclear import of the SMGA-promoter containing plasmids.
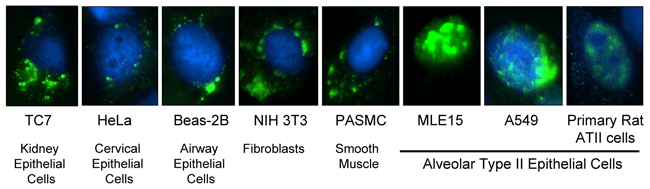
The SPC promoter acts to drive DNA nuclear import specifically in type II alveolar epithelial cells. Plasmids carrying the SPC DTS were microinjected into the cytoplasm of a number of different cell types. Nuclear import of the plasmids was detected in type II alveolar epithelial cells, but in no other cell type.
Moreover, when these factors are knocked down in smooth muscle cells, all nuclear import of the SMGA plasmid is prevented, whereas complexation of the plasmid with purified SRF and Nkx transcription factors allows the protein-DNA complex to now be imported into the nuclei of any cell type. Taken together, these results provide proof of principle for the development of cell-specific non-viral vectors for any desired cell type. Current studies involve expanding our repertoire of cell-specific DNA nuclear targeting sequences so that we may be able to target genes selectively to any desired cell or tissue type. To date, we have identified DNA sequences that support specific nuclear import in osteoblasts, endothelial cells, smooth muscle cells, and type I and type II alveolar epithelial cells.
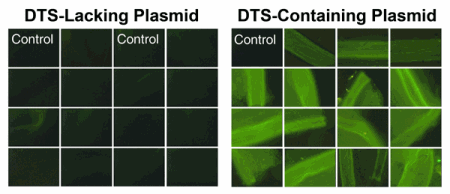
DNA nuclear targeting sequences function in vivo. Plasmids lacking or carrying the SV40 DTS were delivered to mesenteric blood vessels in a number of rats using electroporation and two days later, gene expression was detected. When transfected into dividing cells in culture, these two plasmids express the same levels of GFP, but when delivered to non-dividing cells in these animals, only plasmids carrying the SV40 DTS give significant levels of gene expression.
Most exciting is the fact that these sequences act in living animals as they do in cultured cells. In studies of vascular gene transfer, plasmids containing the SV40 DTS increased gene expression 40 to 200 fold after electroporation-mediated transfer into the mesenteric vessels of living rats. In situ hybridization verified that the increased reporter gene expression in rats electroporated with constructs containing the SV40 DTS vs. constructs lacking this sequence was a result of increased nuclear import of the DTS plasmids. Cell-specific DNA nuclear import sequences also show cell-specificity in vivo. For example, the SMGA DTS was shown to drive nuclear accumulation of plasmids (by in situ hybridization) and enhanced gene expression that was restricted to the smooth muscle cell layer of vessels electroporated with reporter constructs, again proving the utility of cell-specific DTSs for in vivo gene transfer. We have also seen similar restriction of nuclear localization and gene expression to the smooth muscle layer in airways following electroporation- mediated delivery of SMGA DTS-containing plasmids to the lung. These data provide compelling evidence that constructs containing DTSs are capable of driving cell-specific and enhanced expression of transgenes in living animals.




Thank you! It's clear it's not an experiment but an extermination plan:
The best way to have a real dialogue about vaccines being weaponized to handicap, infertilize and murder the “over-population” is to start with vaccine contamination: nobody could be in favor of contaminated pharmaceuticals.
1. Carcinogen SV40 in Oral Polio Vaccine: they knew it since the 60s but kept distributing it even until 2016 !!!
2. hCG in vaccines to infertilize women detected since the 90s: still going on
3. Thimerosal, aluminum, Mono-sodium Glutamate (MSG) and other NEUROTOXINS
4. Heavy metals
5. Human DNA 2000% in excess of FDA 10 ng limit (main driver towards brain damage like autism/asperger/ticks, leukemia and non-Hodgkin cancer), probably related to point 7 below.
6. Graphene oxide in Flu and COVID shots but now with anything injectable (even dentist anesthesia, hospital IV, etc.).
7. Carcinogenic SV40 genomic sequences and double-stranded DNA in mRNA COVID shots: the hacked DNA in the cell doesn’t stop producing the poison when the cell dies, but its descent continue the poisoning until the haccinated casualty dies.
8. Bluetooth nano-routers injected with COVID vaccines and inserted with swabs (which explains why they rejected the cheaper non-invasive saliva test).
Proof of criminal intent:
Points 7 and 8
Censoring and blocking 30+ COVID cures
Labeling the most lethal batches with a lethal code (howbad.info)
Blocking the real knowledge of effectiveness v. "adverse event" rate
That proves:
A. There's zero Government control
B. There's zero Manufacturer liability
C. There's zero Media coverage
D. All that, during decades and still going on, not only with vaccines but also with medicines, food&beverage additives, etc. Everything, even institutions have been weaponized!
E. There's zero political action to stop that (except RFK2 in the USA)
A school buddy told me "I know you make sense but if I recognize it's true, I won't be able to enjoy life anymore".
16 laws we need to exit Extermination Planet
https://scientificprogress.substack.com/p/laws-to-exit-planet-prison
If we don’t succeed, they’ll succeed with their 6-sword lethal plan fully exposed here:
https://scientificprogress.substack.com/p/the-plan-revealed
Change goes in hand with the number of awakened! Thank you for sharing this to save lives!
With the mid September rollout of the Covid Booster and RSV Vax, tens of millions brainwashed Americans lives will be in peril. Warning them is fruitless. I’ve tried. This is exasperating!